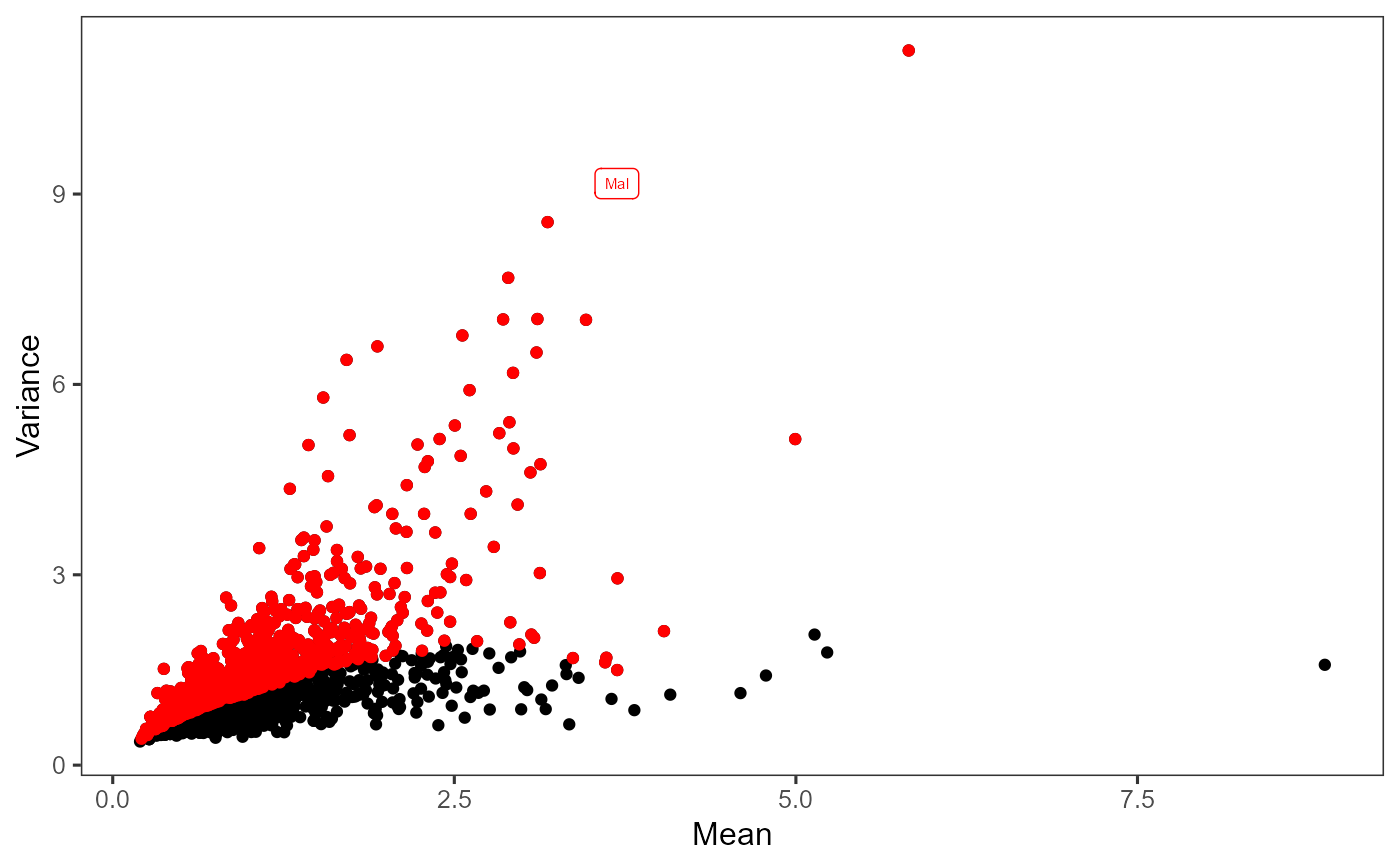
Plot highly variable genes
plotTopHVG( inSCE, method = c("vst", "mean.var.plot", "dispersion", "modelGeneVar"), hvgList = NULL, n = NULL, labelsCount = NULL )
Arguments
| inSCE | Input |
|---|---|
| method | Select either "vst", "mean.var.plot", "dispersion" or "modelGeneVar". |
| hvgList | Character vector indicating the labels of highly variable genes. |
| n | Specify the number of top genes to highlight in red. If
|
| labelsCount | Specify the number of data points/genes to label. By default, all top genes will be labeled. |
Value
plot object
Examples
data("mouseBrainSubsetSCE", package = "singleCellTK") mouseBrainSubsetSCE <- scranModelGeneVar(mouseBrainSubsetSCE, "logcounts")#> Warning: collapsing to unique 'x' valuesplotTopHVG(mouseBrainSubsetSCE, method = "modelGeneVar", n = 1000, labelsCount = 0)